Pca column share how to#
we will use sklearn, seaborn, and bioinfokit (v2.0.2 or later) packagesįor PCA and visualization (check how to install Python packages).For example, in RNA-seqĮxperiments PCA helps to understand the gene expression patterns and biological variation in a high-dimensional Leads to the generation of high-dimensional datasets (a few hundred to thousands of samples). PCA is a useful method in the Bioinformatics field, where high-throughput sequencing experiments (e.g.These top first 2 or 3 PCs can be plotted easily and summarize and the features of all original 10 variables. (generally first 3 PCs but can be more) contribute most of the variance present in the the original high-dimensionalĭataset. PCs are ordered which means that the first few PCs Variables (PCs) with top PCs having the highest variation. (you may have to do 45 pairwise comparisons to interpret dataset effectively). For example, when datasets contain 10 variables (10D), it is arduous to visualize them at the same time.T-SNE can be used for dimensionality reduction for nonlinear datasets. PCA works better in revealing linear patterns in high-dimensional data but has limitations with the nonlinear dataset.PCA preserves the global data structure by forming well-separated clusters but can fail to preserve the.PCA helps to assess which original samples are similar and different from each other.

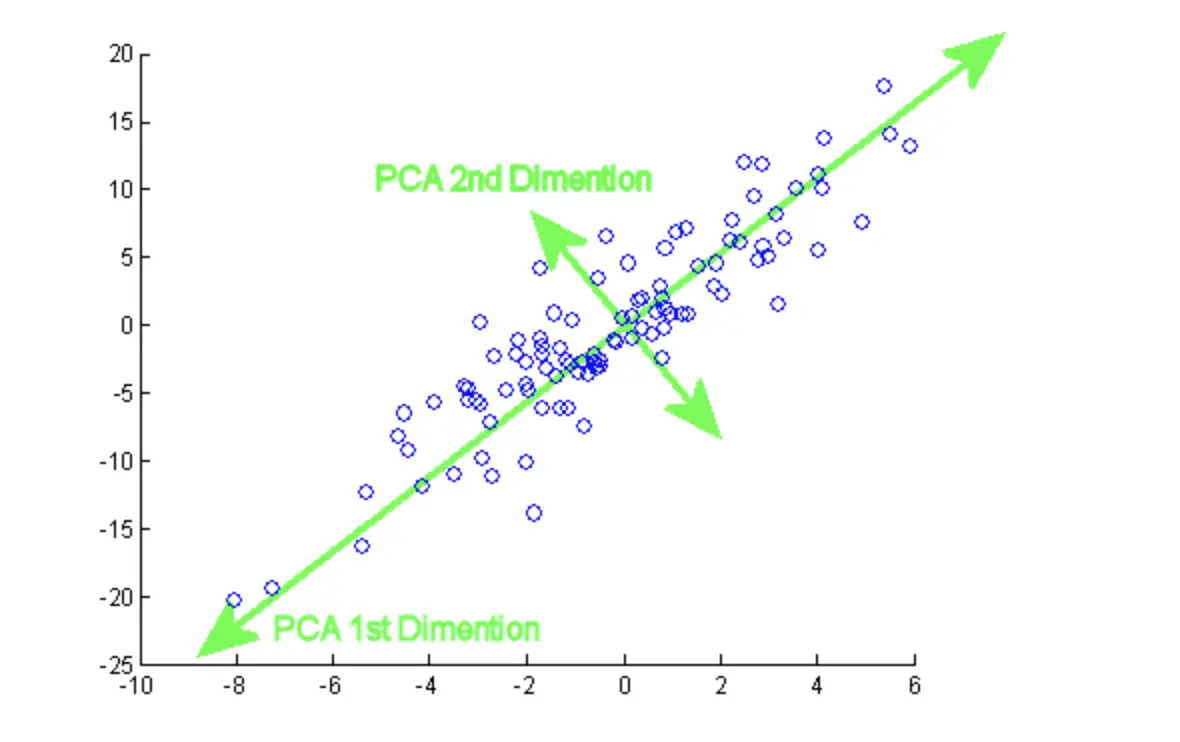
Most of the variation, which is easy to visualize and summarise the feature of original high-dimensional datasets in The first component has the largest variance followed by the second component and so on.New set of uncorrelated variables called principal component (PC) while retaining the most possible variation. PCA reduces the high-dimensional interrelated data to low-dimension by linearly transforming the old variable into a.Method that used to interpret the variation in high-dimensional interrelated dataset (dataset with a large number of variables)